Welcome to the ENETWILD Newsletter!
ENETWILD is an international network of wildlife professionals focused on integrating wildlife management with pathogens’ surveillance and management. The project is funded by EFSA.
In this newsletter you will stay in the loop on the latest publications and updates in the wildlife world, you will have access to event information and will get to know more about the people involved in the project.

Latest publications
🔎 MAMMALNET – Citizen Science Data Collection from a
One Health Perspective ⬇️
https://www.cabidigitallibrary.org/doi/full/10.1079/onehealthcases.2023.0021
🐮 Bird flu outbreak in US cows: why scientists are concerned ⬇️ https://www.nature.com/articles/d41586-024-01036-1
🦠 Epidemiological analysis of African swine fever in the European
Union during 2023 ⬇️
https://www.efsa.europa.eu/en/efsajournal/pub/8809
Events & Useful information
The call of the first EUPAHW (European Partnership on Animal Health and Welfare) external call is now available here and on the new EUPAHW website. You will find the slides of the presentation and useful information at the website.
Topic 1: Novel Technologies for Prevention, Detection, Assessment, and Management of Animal Health and Welfare
Topic 2: Fundamental Research for Animal Health and Welfare
Topic 3: Animal Health and Welfare and Society
Deadline for submissions of pre-proposals is 8th July, 15:00 CEST.

7th European Congress of Conservation Biology
“Biodiversity positive by 2030”: This theme presents a positive message and a call to action towards the conservation of our planet’s biodiversity.
Call for abstract is closed but registration is still open

9–13 September Hybrid format – Stralsund, Germany
15th European Wildlife Disease Conference
“One Health – Challenges and Opportunities for the Surveillance and Management of Wildlife”
The conference will be organized in hybrid format
Data Collection

The creation of an inventory of wildlife population monitoring programmes in Europe is critical to identify knowledge gaps and constraints, as well as research needs and priorities, for an international harmonized approach.
The EUP AH&W will create an inventory, describe, and evaluate wildlife population monitoring programmes and data taking place in Europe. This will be the basis for identifying the limitations, needs, critical knowledge gaps and constraints in existing wildlife population monitoring programs in Europe. This activity will be based on a questionnaire focused on wildlife monitoring systems in EU MS and other European countries, including regional, national (also regional) and international wildlife monitoring systems in Europe.
The questionnaire needs to be returned by 15/06/2024. More information can be found at the ENETWILD webpage.
Updates
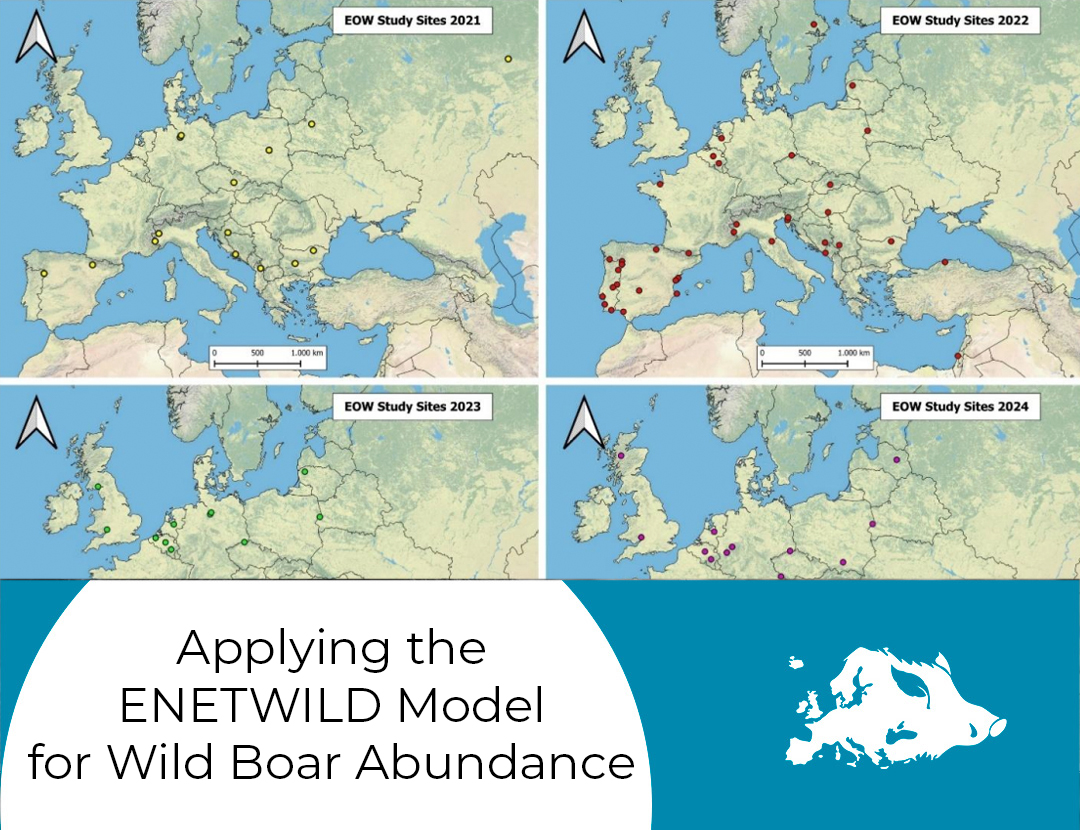
The first training course of the EOW took place on April 10th, with the participation of about 60 collaborators, with many new ones taking part in it, to present activities to new members of the network and start the data collection campaign for 2024.
The main topics of the course were the field protocol for the implementation of the Random Encounter Model and the use of Photogrammetry for the estimation of the parameters directly from the image sequences.
An overview about data processing and analysis was also made, but these topics are going to be explained in detail in the second part of the course, which is going to be held in September.
The 2024 campaign, which is about to start, is expected to include about 58 study sites from 28 countries, reaching the highest numbers the EOW ever obtained.
You can watch the course here:
Interview
Relax and enjoy this interview with Marcus Rowcliffe from The Zoological Society of London

Are you familiar with the ENETWILD project? Why do you think such a project is relevant from One Health’s point of view?
I’ve been working with the ENETWILD project for about two years now. So I’ve not been involved from the start. I’ve come in as a technical specialist, on the monitoring side, but I’m becoming more familiar with it over that time. I think, as we saw with COVID and avian flu threats, and a lot of livestock threats as well, one of the major environmental challenges facing us is the emergence of zoonotic diseases. The One Health agenda is all about creating a holistic view to try to understand tese threats. And I think ENETWILD is a really important project in that space, trying to get a handle on the wildlife abundance side of that.
I think, One Health approaches to understanding zoonotic disease have often done a great job on identifying and monitoring the pathogens, but we generally don’t get such a good picture of the dynamics of the wildlife hosts. So I think this is where this project has the potential to make a really big difference.
Wildlife monitoring can be a more complicated task than it seems. How has your research on wildlife population monitoring helped to improve wildlife management and conservation?
I spend a lot of my time trying to solve technical issues in a way that help us to get good population estimates for wildlife. I don’t always then have a direct connection with work that uses these methods to solve practical problems in the real world. That’s why it’s so exciting to be involved in this project because there is a very direct pathway to the application of methods. What I’m trying to do is develop new statistical methods, and also the tools that go with them to make those methods accessible, making it easier and more straightforward to get good, reliable abundance estimates for wildlife. So, I mentioned that a gap in the One Health agenda has been getting really good estimates of wildlife dynamics. The methods that I’m trying to develop, are intended to plug that gap. So, over time, we start to build that picture of dynamics that will help us understand and manage the health risks. And having accessible, easy-to-apply methods is really crucial for that.
Can you give us a brief overview of what the Random Encounter Model (REM) entails and how it works? (Widely used at ENETWILD and EOW)
The idea with the random encounter model is to get animal abundance estimates from camera trapping data. We’ve known ever since camera traps were used, that there is a very rough correlation between the rate at which you get wildlife records from camera traps and the abundance of that wildlife. But there are many other factors that can influence that relationship. If your cameras are more effective, they can see further, the vegetation is less dense, temperature is different, and so on and so forth, lots of different things can affect trap rate as well as abundance. What the random encounter model does is, to borrow from the physical gas model theory, to model the pattern of contacts between moving particles, as an analogy for animals and static articles cameras. Using a process model of this kind allows us to control for the confounding factors that may also influence the trap rate as well as abundance, captured as the detection zone dimensions and the animal speed. And we can now measure those key variables from the camera data by tracking animal positions as they pass the camera, using computer vision techniques.
How does the REM differ from traditional methods of estimating population densities of wildlife?
I guess it depends on what you mean by traditional. The really old-school traditional approach would be trying to count animals directly in the field, so going out with a notebook and counting what you can see. The big difference there is that, while cameras are mostly ignored by animals, direct observation methods just don’t work well for the kinds of animals we’re trying to count as they tend to be very wary of humans, so it’s often impossible to get reliable abundance estimates that way. There are also long-established methods based on sign surveys (like dung counts), but they have a lot of problems around understanding the different factors that can confound the relationship between sign density and animal density, so they can be useful but are not perfect. In terms of existing camera trap methods, the first (in a sense traditional) wildlife abundance estimates from camera traps were based on individual recognition and mark recapture methods, but these obviously required that the animals has some kind of individual marking, like leopards or tigers. But working with wild boar, for example – a key EOW target species – we can’t reliably identify sufficient individuals to apply those methods. The REM is one of a suite of related methods (including distance sampling and time to event models) that enables us to estimate the abundance of unmarked species from camera trap data.
Could you share some notable successes or applications of the REM in real-world conservation or research projects?
There are a few examples where it has been possible to make the first abundance estimates for some highly threatened species, for example some endemic deer and pig species of Bawean Island in Indonesia. The species have very small ranges, so REM abundance estimates have provided a good idea of the global population sizes – only a few hundred individuals in both cases. This has provided importance evidence for IUCN threat assessments. Similarly, I’m advising a research student currently working on armadillo species in Brazil, particularly a narrow range habitat specialist species about whose ecology we know very little at the moment, including basic information on typical abundance patterns. An example related to EOW goals is a project I’m involved with in the UK, where we are evaluating the vaccination of free-living badgers as a possible way to limit or eliminate their role in the transmission of bovine tuberculosis in the UK. We are currently testing whether the REM can help us to track the effectiveness of vaccination efforts by estimating local population size and hence vaccination coverage. The preliminary results of this work are look like the approach has potential.
The EOW is using some of the methodologies and tools you have developed. What advantages do you think initiatives like the EOW have for wildlife managers?
I think the main potential benefit is that you have a consistent methodology that can be shared across teams and it can give you robust repeatable abundance estimates which will then build over time to give you an idea of wildlife dynamics.
What are some of the limitations or challenges associated with implementing the REM, and how do you suggest overcoming them?
The big challenge is the size of the technical task in processing imagery, generating analysable data from that and having a smooth pipeline all the way from image through to ecological information. It’s actually a huge task because camera traps generate enormous amounts of imagery, and we have to say what’s in those images. For the REM, we also have to measure how far animals are from the camera. As you would expect, we’re using new image processing technology to try to reduce the human time required for processing as far as possible. Methods are getting better in this area at a very rapid rate and so what we’re trying to do is to keep on top of developments in order to harness the best of them to apply in this particular challenge, developing models that not only work very well for image recognition and depth perception but are then built into tools that are accessible and easy for non-specialists to use. The combination of cutting-edge models with highly accessible tools has been mostly lacking till now, and it’s something that we’re trying to really get on top of in the EOW project. There’s lots of work still to do there and lots of improvements to emerge, but things are moving forward.
You mentioned that with the camera traps you have a lot of images and a lot of data to process and analyse, so I’m sure you know of citizen science initiatives that make accessible tools and that can help with this part, where people that are non-specialized could help in this field?
EOW is not using this approach at the moment. We’re using a combination of computing tools and a relatively small number of specialist users who can guide that process. But there are indeed some platforms that focus on mass participation from citizen scientists in image processing. I’m involved in another project focused on national wildlife monitoring in the UK which is trialling that approach, where we are seeking to find the optimal combination of machine, citizen and specialist input to minimise human time requirements while keeping the accuracy of processing high. In this project we also recognize that engagement, outreach and public understanding of science are benefits of the citizen science approach in its own right.
How important is it for conservationists and researchers to accurately estimate population densities of unmarked animals, and how does the REM contribute to this goal?
There’s a whole range of ways in which animal abundance information can be useful, particularly in the one health context where we’re trying to understand the dynamics of pathogens and how wildlife host reservoirs contributes to that. Fluctuations in the abundance of wildlife can be a key driver of the potential for pathogens to spill over into human or livestock populations, so being able to monitor those fluctuations is really crucial there. It can also help us to answer specific questions in the one health space, like the vaccination coverage issue I mentioned earlier. It’s also essential if we’re trying to conserve highly threatened species like the ones mentioned earlier. If we don’t know what’s happening to those populations, then it’s very difficult to react appropriately and prioritize conservation actions for them. It also can be extremely useful to have an idea of the absolute abundance of hunted populations, so we know what a sustainable harvest might look like. So lots of reasons that we might want to get accurate abundance estimates of animal populations. The REM approach contributes to these goals because many of the species we’re talking about are hard to monitor any other way.
Finally, what advice would you give to aspiring conservationists, wildlife researchers or is there anything you would like to highlight for the readers of the newsletter?
I think my top piece of advice for people starting out in this field is to be as enthusiastic as you can and meet as many people as you can. Seize every opportunity that comes your way. If you see meetings that are coming up which you’re interested in, try and get along to those and talk to people there, look out for people advertising positions like research assistantships or working on practical conservation projects, even if they’re short term. That puts you in a good position to build contacts, build experience, shows that you’re strongly focused on succeeding in this area, and it should also be a lot of fun having that diversity of experience. Just get out there and pick up as many opportunities as you can.
We need your help!

The iMammalia recording app has reached nearly 25,000 mammal records across Europe since 2019. It is currently available in 17 languages, and we are looking for volunteers to complete translations into Bulgarian, Hungarian, Romanian and Lithuanian in particular, to increase the number of records in Eastern Europe.
If you know of someone who may be interested, please get them to contact Graham Smith at graham.smith@apha.gov.uk.
New website!
Go and visit our brand-new website, now there are some new sections such as:
- Mission & Objectives
- Workpackages
- Direct links to EFSA’s ASFStop campaign
- New section on ONE HEALTH, including its background and results